Anne Nakamoto
Bioinformatics PhD Candidate at UC Santa Cruz
Research
California Conservation Genomics Project
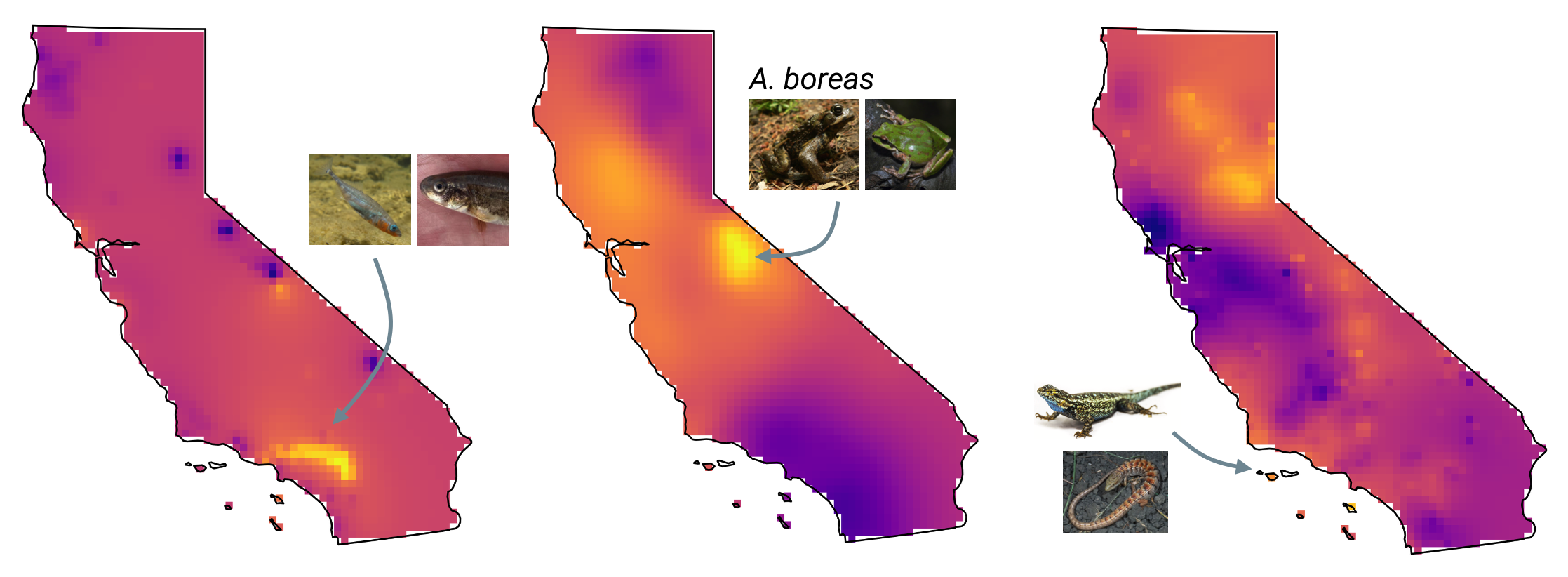
I perform analyses investigating population health for the California Conservation Genomics Project (CCGP). In particular, I study differences in mutation load between CCGP species populations across the landscape of California.
Evolutionary histories of invasive fungal pathogens causing rapid death of the native ‘ōhi‘a tree in Hawai‘i
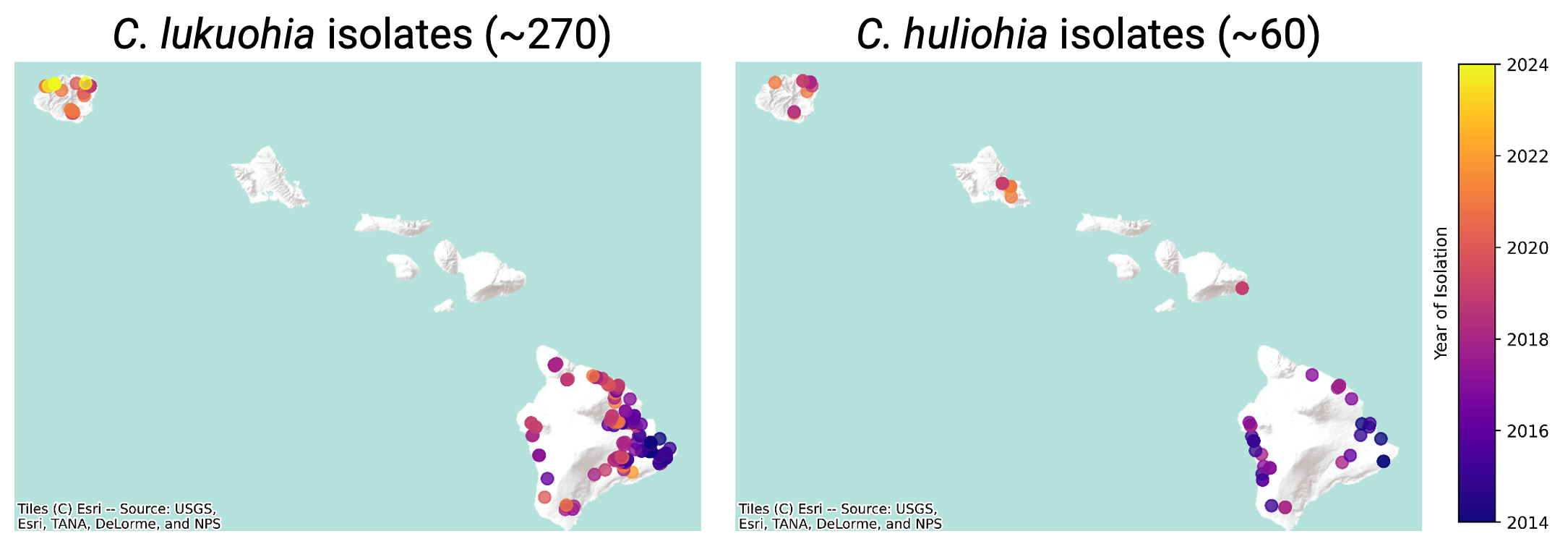
Hawai‘i’s native forests are threatened by Rapid ‘Ōhi‘a Death (ROD), a disease of the keystone and culturally significant native ‘ōhi‘a tree (Metrosideros polymorpha) species. ROD was first characterized in 2014, and is caused by two novel, non-native fungal pathogen species, Certaocystis lukuohia and huliohia. Previous work has shown that C. lukuohia causes more aggressive disease, and likely originated from a single introduction to Hawai‘i Island within the past few decades. Meanwhile, C. huliohia is less aggressive, contains greater genetic diversity, and may have been introduced much earlier. Given these differences, there is currently much discussion regarding the respective management strategies of the two ROD pathogens. However, the evolutionary histories and invasion dynamics of both pathogens are not well understood, and their genomes remain largely uncharacterized.
Anne Nakamoto, Lisa Keith, Qingyi Yu, Lionel Sugiyama, Xiaohua Wu, Blaine Luiz, MaryAnn Villalun, Jodie Jacobs, Russ Corbett-Detig, Ariana Cisneros, Harrison D. Heath, Cole Shanks, Faith Okamoto, Alexis Abigail Aroma Albura, Kyle Henricson, Yi Jun Lan, Henry Moore, William Seligmann, Yulia Zybina. Genome sequence of Ceratocystis huliohia, a fungal pathogen of the native ‘ōhi‘a tree in Hawai‘i. In review at Microbiology Resource Announcements. github
Transposable element dynamics
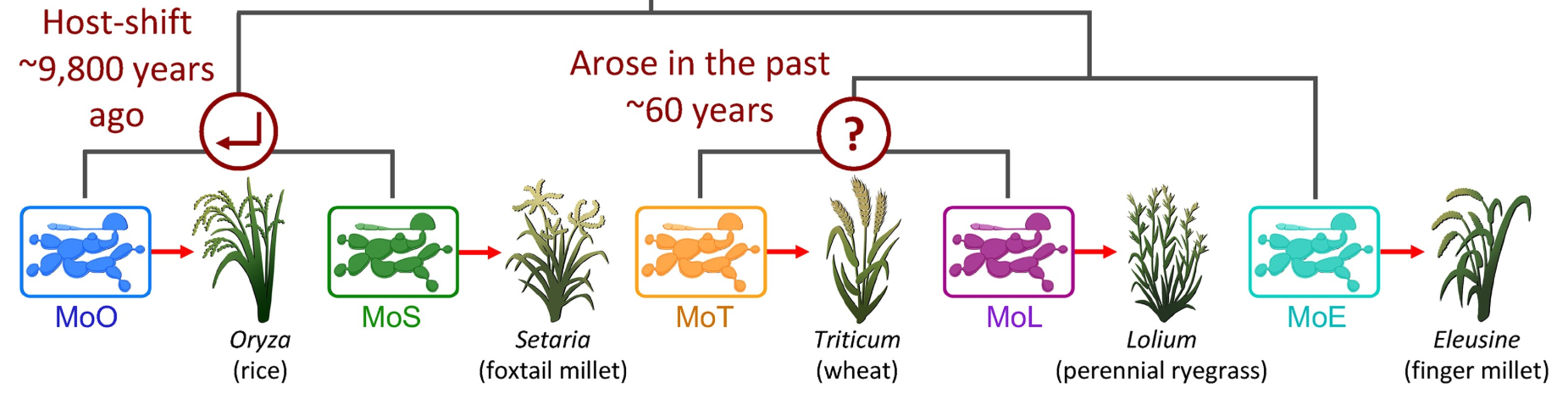
Anne Nakamoto,* Pierre Joubert,* Ksenia Krasileva. Intraspecific variation of transposable elements reveals differences in the evolutionary history of fungal phytopathogen pathotypes. Genome Biology and Evolution, 15 (12), evad206. (*authors contributed equally)
Landen Gozashti, Anne Nakamoto, Shelbi Russell, Russell Corbett-Detig. Horizontal transmission of functionally diverse transposons is a major source of new introns. Proc. Natl. Acad. Sci., 122 (21) e2414761122.
Genomic Resources
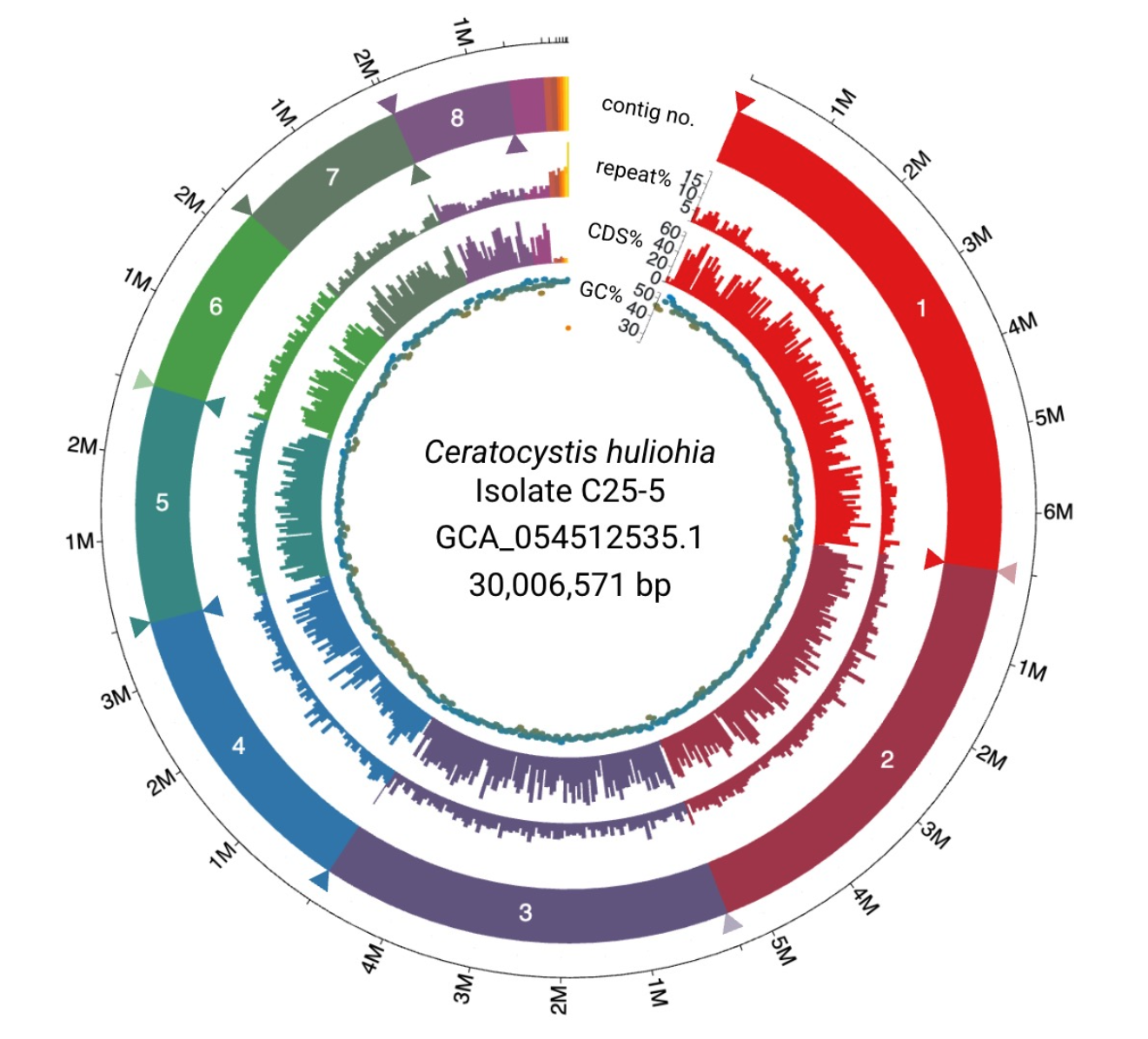
I am currently leading the generation of long-read genome assemblies for Ceratocystis lukuohia and huliohia, to facilitate comparative genomic study of the Rapid ‘Ōhi‘a Death disease. The first of these genomes for C. huliohia is available on NCBI. More to come!
Teaching
BMEB Bootcamp for 1st year graduate students
I was an instructor for the BMEB Bootcamp, a two-week orientation for incoming PhD and Masters students led by the T32 Genome Sciences Fellows in the Department of Biomolecular Engineering & Bioinformaticcs at UC Santa Cruz. I led hands-on genome sequencing and assembly tutorials.
Genomes and Plant Health Workshop
I was an instructor for the Genomes and Plant Health Workshop, hosted by Ksenia Krasileva's Lab in the Department of Plant and Microbial Biology at UC Berkeley. I taught undergraduate and high school students a section on genomics and computational biology, and developed a Google Colab tutorial to teach transposable element (TE) annotation.
